- The Utrecht Biomolecular Interactions software portal provides access to software tools developed in the Computational Structural Biology group / NMR Research Group of Utrecht University with a main focus on the characterization of biomolecular interactions. Please note that this site is in active development.
- HADDOCK Web Docking
 HADDOCK (High Ambiguity Driven protein-protein DOCKing)
is an information-driven flexible docking approach for the modeling of biomolecular complexes. HADDOCK distinguishes itself from ab-initio docking methods
in the fact that it encodes information from identified or predicted protein interfaces in ambiguous interaction restraints (AIRs) to drive the docking process.
HADDOCK can deal with a large class of modelling problems including
protein-protein, protein-nucleic acids and protein-ligand complexes. | Go to
service >>
HADDOCK (High Ambiguity Driven protein-protein DOCKing)
is an information-driven flexible docking approach for the modeling of biomolecular complexes. HADDOCK distinguishes itself from ab-initio docking methods
in the fact that it encodes information from identified or predicted protein interfaces in ambiguous interaction restraints (AIRs) to drive the docking process.
HADDOCK can deal with a large class of modelling problems including
protein-protein, protein-nucleic acids and protein-ligand complexes. | Go to
service >>
- PRODIGY
- PRODIGY (PROtein binDIng enerGY prediction) webserver predict of the binding affinity in protein-protein complexes. To use PRODIGY you just need to provide the three-dimensional structure of your complex/complexes as PDB/mmCIF format. | Go to service >>
- DISVIS
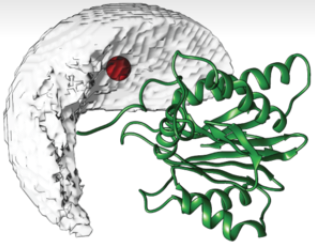 DISVIS allows you to quantify and visualize the information content of distance restraints between macromolecular complexes. | Go to service >>
DISVIS allows you to quantify and visualize the information content of distance restraints between macromolecular complexes. | Go to service >>
- POWERFIT
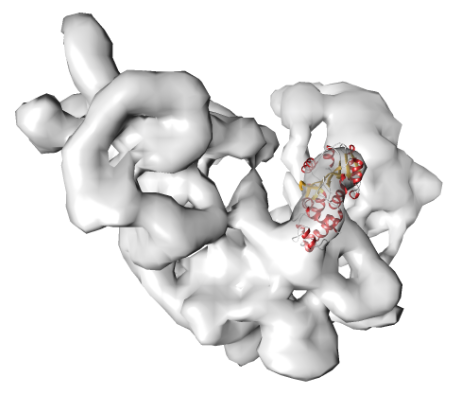 POWERFIT allows you to automaticall fit atomic models into cryo-EM density maps. | Go to service >>
POWERFIT allows you to automaticall fit atomic models into cryo-EM density maps. | Go to service >>
- CPORT
- CPORT is an algorithm for the prediction of protein-protein interface residues. It combines six interface prediction methods into a consensus predictor. CPORT predictions can be used as active and passive residues in HADDOCK. | Go to service >>
- WHISCY, Protein-Protein Interface Prediction
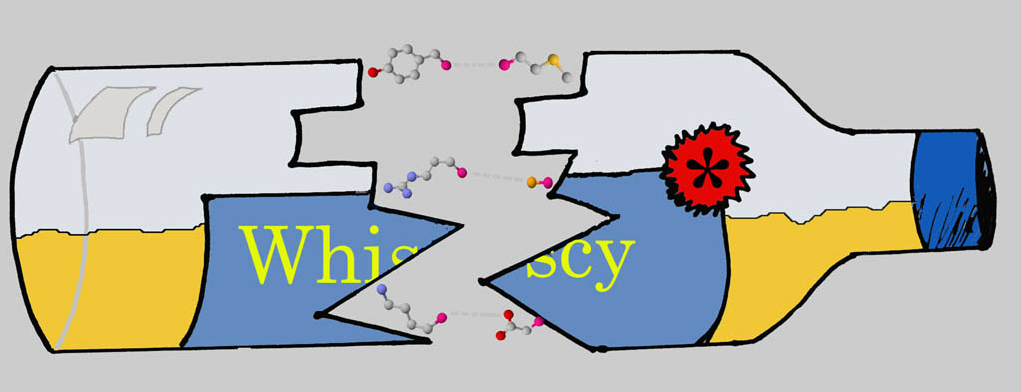 WHat Information Does Surface Conservation Yield? WHISCY is a program to predict protein-protein interfaces. It is primarily based on
conservation, but it also takes into account structural information. A sequence alignment is used to calculate a prediction score for each surface residue of your
protein. | Go to service >>
WHat Information Does Surface Conservation Yield? WHISCY is a program to predict protein-protein interfaces. It is primarily based on
conservation, but it also takes into account structural information. A sequence alignment is used to calculate a prediction score for each surface residue of your
protein. | Go to service >>
- FANDAS
- FANDAS 2.0: Fast Analysis of multidimensional NMR DAta Sets (FANDAS) is a tool to predict peaks for multidimensional NMR experiments on proteins. | Go to service >>
- PROABC-2
- proABC-2 predicts which residues of an antibody are forming intermolecular contacts with its cognate antigen. | Go to service >>
- FCC CLUSTERING
- RMSD-based clustering of complexes can be slow and is inadequate for large multi-molecular complexes, particularly when their components are symmetric. We developed a novel clustering strategy that is based on a very efficient similarity measure - the fraction of common contacts. | Read more>>






